|
5/6/2023 0 Comments Alpha helixThe sliders at the bottom of the graphics window work just like ordinary scroll bars: you can move them slowly by clicking on the arrows, or make larger jumps by clicking in the open scrollbar, or you can drag the slider. Try out the ZOOM function, starting from View1. Check to see if this alpha helix has 3.6 residues per turn. Click on backbone atoms at either end of one of the H-bonds, to verify that the alpha-helical H-bond pattern does indeed go from a donor NH at residue i to an acceptor O at residue i-4. Turn on "Hbonds" on the button panel, to see the H-bonds in brown. 
Lets use this tool to explore this helix- how long is the helix? What is the diameter of the helix?Ĭhoose View4 for a close-up from the side, with the helical hydrogen bonds (H-bonds) in brown. You can measure the distance between two atoms, or between the same atom in two different conformations. Pick a second atom - the pointID for the second atom is shown, and the distance between the two atoms is given. A 'pointID' for that atom is printed at the bottom left of the graphics window this would usually contain residue name and number and atom name, but can be freely specified in the kinemage file to be any character string. Now try out some informational features of KiNG. Grisham (University of Virginia in Charlottesville, Virginia). This figure is a snaphot of a Java Applet written by Edward K. As an ideal alpha helix consists of 3.6 residues per complete turn, the angle between two residues is chosen to be 100 degrees and thus there exists a periodicity after five turns and 18 residues. The amino acid side chains are projected down the axis (the axis of an alpha helix, orthogonal to the paper plane). To reset the image to the start-up view, pull down the 'Views' menu and release on View1.įigure (left): A helical wheel representation of an amino acid sequence. Choose View2 or View3 to see more of the structure, which provides the context to understand the distribution of amino acids along the helix. This is a typical globular-protein helix. 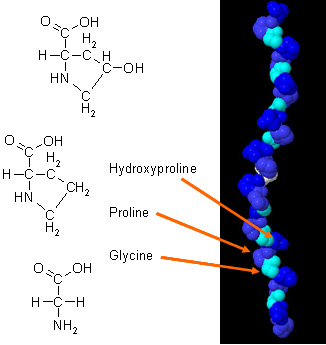
When you clicked the different sidechain types on, what did you observe? Did you notice that the helix has one side facing toward the solvent with mainly polar residues, and the other side toward the protein interior with mainly hydrophobic residues. Turn on and off the 'static' group and the display objects underneath it, to see how the button logic works: an individual object can be turned off either by its own button or by a subgroup or group above it in the heirarchy. Now TURN ON and OFF the various display groups and sets, by clicking in the appropriate button box. These can be turned on by clicking on the checkbox labeled "side ch". The hydrophobic side chains are shown in seagreen, polar ones in skyblue, and charged ones in red. Notice that the Calpha-Cbeta bonds do not point out radially from the helix axis but "pinwheel" along the line of one of the adjacent peptides, giving the side chains an asymmetrical start. The non-integral, 3.6-residue-per-turn repeat of the alpha helix means that the Calphas of successive turns are about halfway offset, giving the main chain a distinctive 7-pointed star appearance in end view. The O and N atoms of the helix main chain are shown as red and blue balls, respectively. This kinemage shows an individual alpha helix, viewed from the N-terminal end to resemble the "helical wheel" (see figure below). Depth-cuing is done primarily by color intensity there is also a simple sort of hidden-line removal. Smoothness of rotation depends on the number of vectors in the display and the speed and size of the computer. Use the mouse to ROTATE the image: dragging across the top of the graphics window will rotate around Z (in the plane of the screen) and dragging anywhere else will rotate around the Y or the X axis or a combination. Upon loading this page, the KiNG Java Applet should automatically spawn. 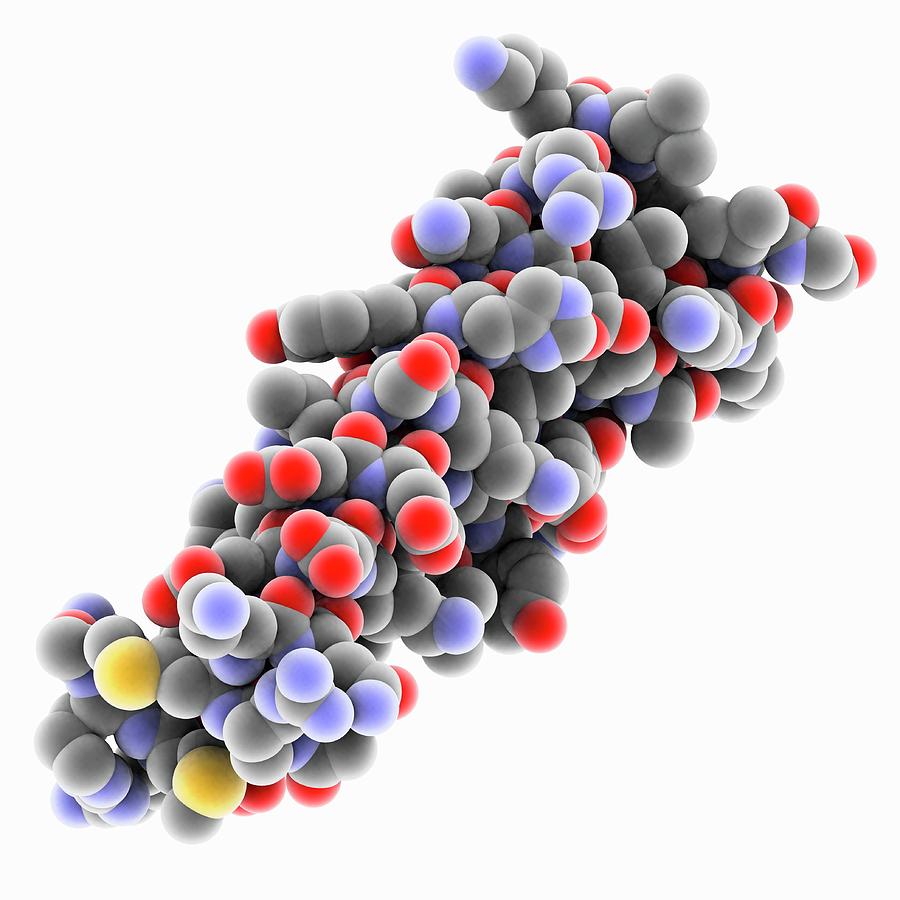
To load the KiNG Java Applet, just click here. The text window is scrollable the graphics image is rescaled when its window is resized. 
The windows can be moved around by dragging their top bars, or resized by dragging in the lower right corner. Detailed suggestions are given below, and should eventually be read to make sure you do not miss any of the major functionalities. Many features of the kinemage display are self-explanatory, and can be worked out by exploring the possibilities yourself. Usually the text window is hidden directly behind the graphics window. KiNG is composed of two windows- a Text window that gives a short description of the molecule being viewed in the larger window.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
